and some more, lol.
What a (beautiful) mess 
1 Like
haha yeah indeed.
Awesome @Nseraf !
I would just be interested in explanations of scientists about what we see 
For example about the cell types, like Ipsi-/Contra-cells (see blog: A few Mystic cells )
Pic 54 for example shows different groups of neurons with different orientations. Can you name them?
What is orientation for example in pic 43 (dorsal,ventral,medial,lateral)?
What sort are the thick, long neurons for example in pics 78,79?
What kind of cell is the beautiful big red-cyan neuron?
Can the original integrator-neurons be seen in the pics (like having their own color)?
@Nik : Just an idea: Can you create a pic showing one integrator neuron and some of the neurons synapsing with it (not the 100 in average as expected) so one can see better the connections? (and add the frame for knowing relative position in dataset). 
1 Like
I can, but I have no way of knowing which of the cells synapse with them.
I’ll have to do some “research”, /add-cell some of the integrator neurons with some of the other cells (some synapse with M-cell btw) and find out which synapse with which integrator neuron, if you want you can help with that 
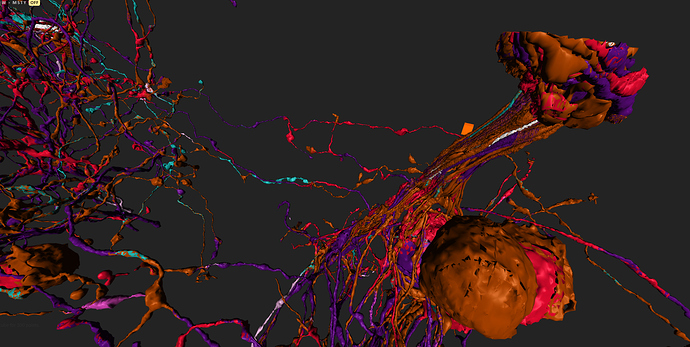
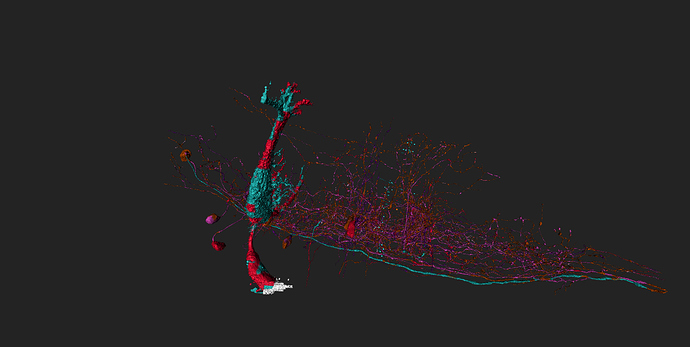
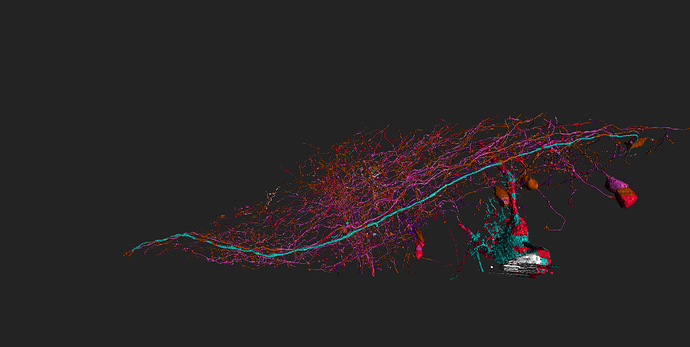
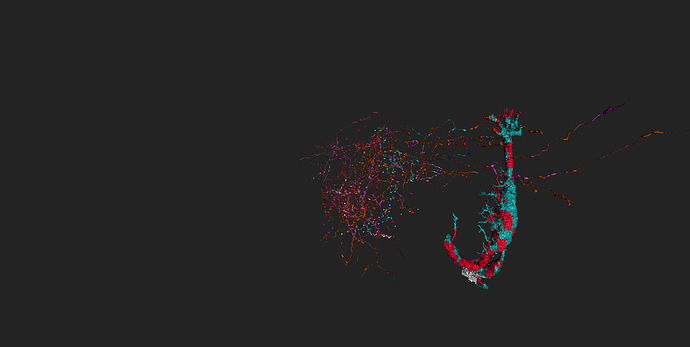
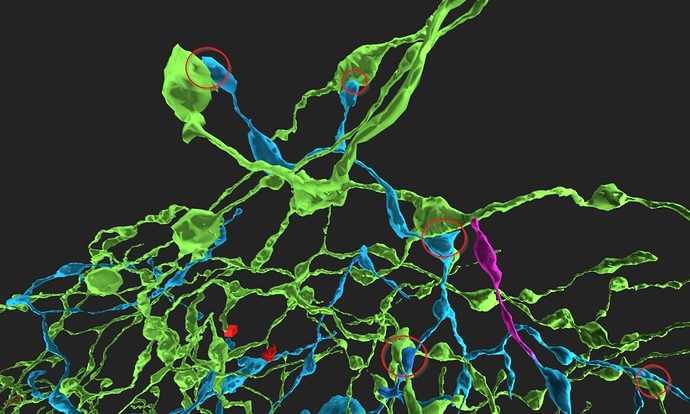
took integrator-neuron 76198 (Old1.1) grass-green
and added neurons synapsing with it (cell-ids 76771 to 76787) cyan
and integrator-neuron 76183 (old 2_2) light blue
and added neurons synapsing with (ids 77856-77873) grass-green
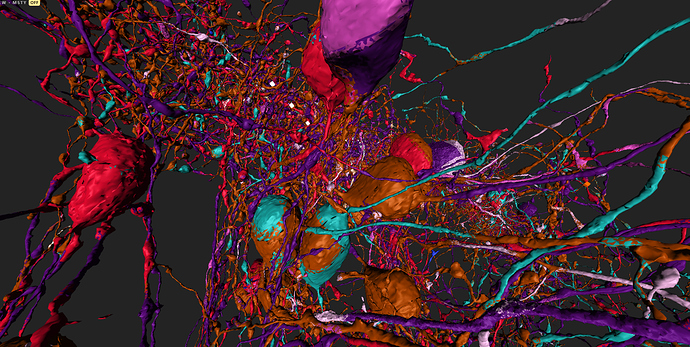
some of (possible) synapses marked
3 Likes
lol, like kk said, what a beautiful mess.
Whoa that’s a lot of cells!
1 Like
Awesome! So many cells! @susi I’m posting this question to the zfish science team. They have an important site visit from out funders going on so it might be a little but before they can respond, but we will make sure they do!
2 Likes
looking forward 
1 Like
These images are awesome @Nseraf! I am not sure how you are generating them, but assigning individual colors to each cell would be cool. To answer some of @susi’s questions
- Pic 54 for example shows different groups of neurons with different orientations. Can you name them…
Unfortunately, not yet, but that is also becase we do not know wht most of the cells in this area look like. You can look at this database to see if there are any similar looking neurons( http://engertlab.fas.harvard.edu/Z-Brain/#/home).- What is orientation for example in pic 43 (dorsal,ventral,medial,lateral)?
the Right of the screen is Rostral (head), Left is caudal (tail). The plane that is in the foreground is medial and in the background is lateral. Bottom is dorsal and top is ventral.
3 What sort are the thick, long neurons for example in pics 78,79?
Usually thick axons mean they are myelinated axons. They tend to travel longer distances, thus they are thicker.
Most of those synapses marked in the following pics are correct, you can look at the 2d plane to see the actual synapse.
4 Likes
i /add-cell (and with crazy’s script I can add more than 1 cell per use of command (to max chars of the msg in chat about 10-20 cells per time I think) i can make abt 50-100 cells per pic before browser crashes lol (thus nearly 90 pics for 1700 cells lol) as to the colours i cant assign colour per cell, i just had scythe vision on and so all cells took those colours depending on the colours of each cells cubes.
You can select None as a heatmap and then each cell will have a separate color. With such big number of cells together, some colors will repeat, but still, they should be easier to differentiate.
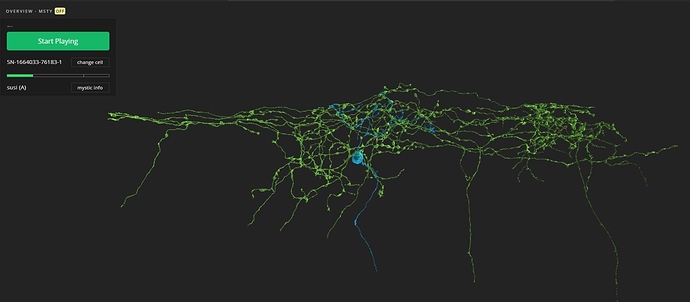
Here’s an example with some random cells taken from The Registry:
There are about 10 cells. Unfortunately the colors already started to repeat. But for a couple of cells, that might work good. I’ll take a look at the EW code to see, if it’s possible, to assign a color to a cell while adding it to the ov. Maybe a /remove-cell command would be useful too.
3 Likes
those commands/ideas sound great! 
Also I saw it with my pics as well, but in your pic it is more apparent/evident. So, I find it interesting that some, at least, zfish cells seem to go diagonally rather than horizontal/vertical in the dataset, I wonder why that is? If it is investigated and so on.
2 Likes
Thanks very much ashwin for your explanations! And for the link, i will take my time to look into it!
If you have any ideas what we and expecially Nseraf could create which helps you, please tell 
2 Likes